Structural Biochemistry/Enzyme Catalytic Mechanism/Enzyme Classification/ Ligases - Wikibooks, open books for an open world

Structural Biochemistry/Enzyme Catalytic Mechanism/Enzyme Classification/ Ligases - Wikibooks, open books for an open world
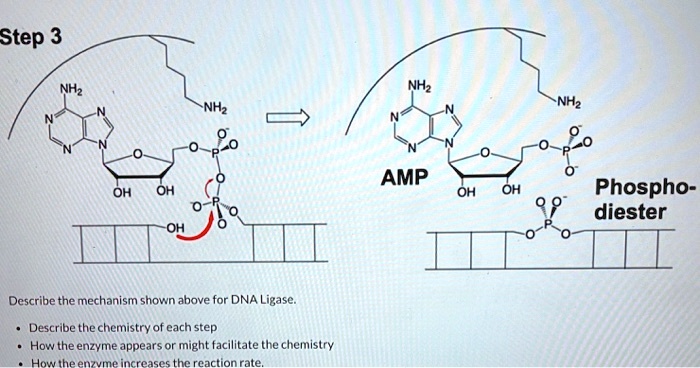
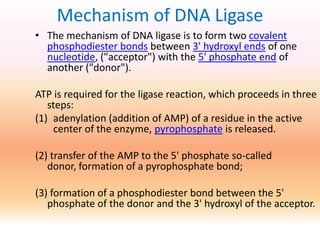
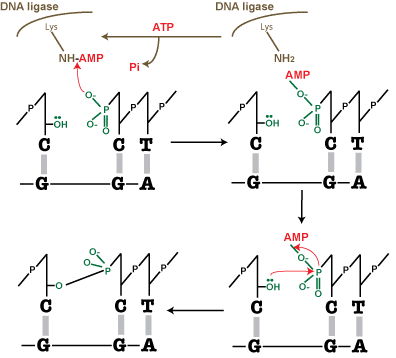
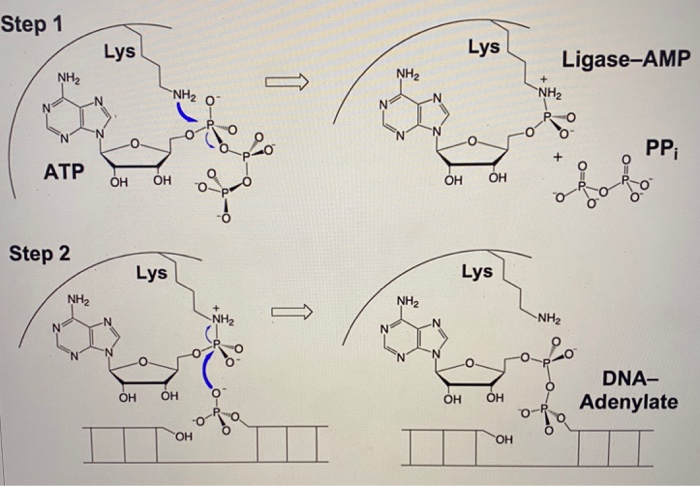
SOLVED:Step 3 AMP OH OH OH OH Phospho- diester Describe the mechanism shown above for DNA Ligase: Describe the chemistry of cach step How the enzyme appears or might facilitate the chemistry

RNA Ligase Structures Reveal the Basis for RNA Specificity and Conformational Changes that Drive Ligation Forward: Cell
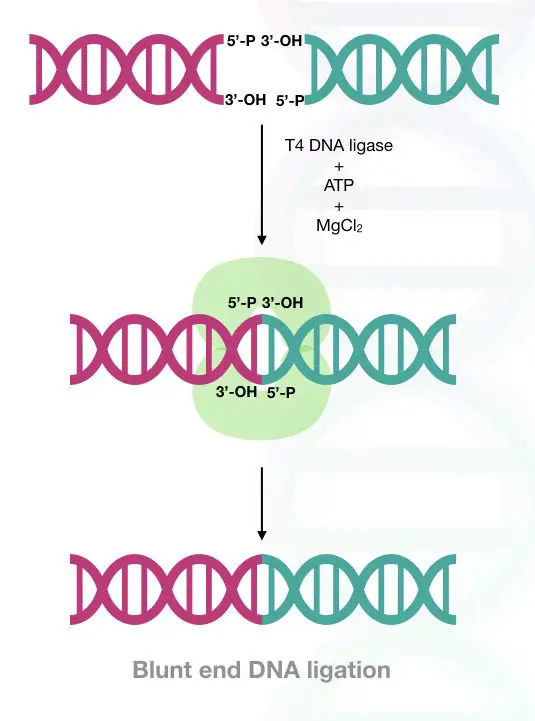
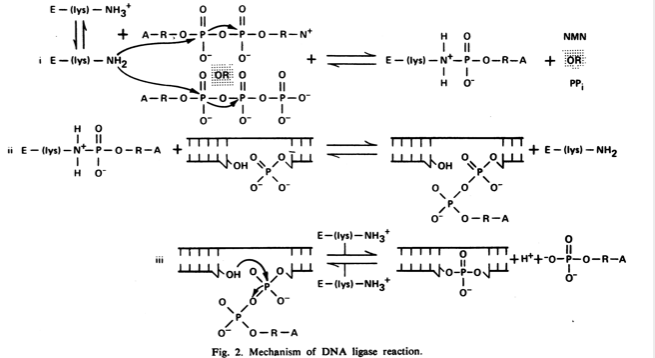
DNA ligation reaction. The reaction mechanism shared by ATPdependent... | Download Scientific Diagram

Structural and mechanistic insights into guanylylation of RNA-splicing ligase RtcB joining RNA between 3′-terminal phosphate and 5′-OH | PNAS

Figure 6 from Human DNA ligase IV and the ligase IV/XRCC4 complex: analysis of nick ligation fidelity. | Semantic Scholar